List of Software and Web Applications
To ensure compatibility and optimal performance, download and
install the latest version of the software that matches your
computer's operating system, unless a specific version is required.
Module 1: Introduction to Linux
| Software or Website |
Operating System |
URL |
| WSL |
Windows |
WSL Installation Guide |
| Default Terminal app |
Mac |
Pre-installed |
| Any terminal or shell |
Linux |
Pre-installed |
Module 2: Next Generation Sequencing
| Software or Website |
Operating System |
URL |
| Java |
All, Install also in WSL |
Download
Java |
| Homebrew |
Mac only |
Homebrew
Website |
| BWA (Burrows-Wheeler Aligner) |
Windows (in WSL), Mac, Linux |
BWA
Website |
| GATK (Genome Analysis ToolKit) |
Windows (in WSL), Mac, Linux |
GATK Website |
| FastP (FastQ Preprocessor) |
Windows (in WSL), Mac, Linux |
FastP
GitHub |
| FastQC (FastQ Quality Control) |
Windows (in WSL), Mac, Linux |
FastQC Website |
| Picard Tools |
Windows (in WSL), Mac, Linux |
Picard Tools Website |
| SAMtools |
Windows (in WSL), Mac, Linux |
SAMtools
Website |
| Bcftools |
Windows (in WSL), Mac, Linux |
Bcftools Website |
| R |
Windows (in WSL), Mac, Linux |
R
Project Website |
| R-Studio |
Windows (in WSL), Mac, Linux |
R-Studio Download |
Module 3: Genome-Wide Association Studies
Module 4: Post-GWAS
Additional instructions specific to a software:
1. Plink
Plink can be downloaded from here.
Download the binary distribution for your system from the
"Binaries" table, "Stable build" column. Then unzip the folder and
move it to the training directory.
Direct links (as of 2024-06-06):
- Linux binary to use in WSL2
- Windows binary to use in cmd.exe /
PowerShell
- MacOS users need Mac binary
Note: PLINK is not a regular app to be installed
into Program Files. It is just a command-line executable. You only
need to download it and make sure you have privileges to execute it
(i.e. admin privileges).
To check installation, open PowerShell, navigate to the folder
containing
plink.exe
and type:
.\plink.exe --help
Alternatively, you can use
cmd
in the address bar:
- From the Plink folder, go to the address bar and highlight
the address.
- Type
cmd in the address bar.
- Type
.\plink.exe --help
Watch a video on how to install on Windows.
2. R
R can be downloaded from here.
For the mirror, use this link.
To install "source-only" packages on Windows:
You'll need Rtools from here.
- If your version of R is R-4.2.xx, install Rtools42.
- If your version of R is R-4.3.xx, install Rtools43.
3. R Packages on R-Studio
Install the following R packages in R-Studio:
install.packages("qqman")install.packages("ggplot2")install.packages(c("dplyr", "glue",
"reshape2"))install.packages(c("readr", "vroom",
"openxlsx"))install.packages("BEDMatrix")install.packages("BiocManager") - To install
packages from BioconductorBiocManager::install("snpStats")install.packages(c("shinydashboard", "DT",
"bslib", "rmarkdown"))
4. Optional Software
The following software are optional, but may be useful:
Post-Installation Check
1. Java
Check the version of Java by opening a PowerShell or Terminal
and typing:
java -version
2. TASSEL
Open TASSEL and use the tutorial dataset packaged within the
Tassel folder.
3. RStudio
Open RStudio, and run the
version
command in the console.
Install the
qqman
package using the menu.
4. Powershell/Terminal
Open PowerShell (Windows) or Terminal (Mac) and note the current
working directory by typing
pwd
then pressing Enter.
5. WSL
If you are on Windows, check the WSL version:
wsl --version
wsl --list
Software Installation Guide
WSL2
Helpful links:
Make sure Windows is up to date. It is recommended to run
Windows 10 version 2004 and higher (Build 1904 and higher) or Windows
11.
- Go to Turn Windows feature on or off in the Control
Panel.
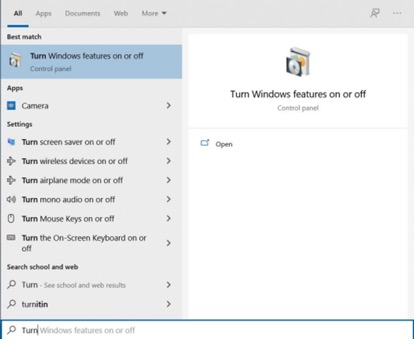
- Enable Windows Subsystem for Linux and Virtual
Machine Platform.
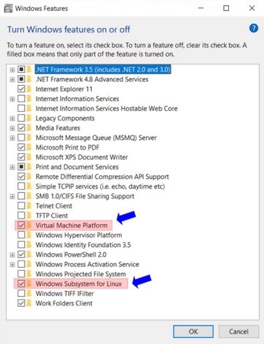
- Restart the computer.
- Download the latest WSL2 kernel here.
- Follow the instructions in the link for installation.
Ubuntu Installation
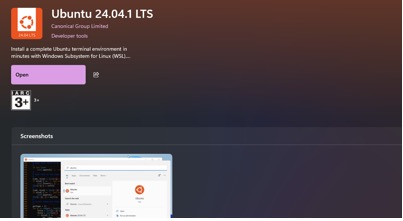
- Go to the Microsoft store and search for Ubuntu.
- Install Ubuntu 24.04.1 LTS.
Software (Install in Ubuntu)
FastQC
sudo apt-get install fastqc
FastP
sudo apt-get install fastp
BWA
sudo apt-get install bwa
SAMtools
sudo apt-get install samtools
BCFtools
sudo apt install bcftools
JAVA Installation
Download the latest package (JDK24) for Linux / MacOS or use
the following command:
wget https://download.oracle.com/java/24/latest/jdk-24_linux-x64_bin.tar.gz
Place the file in your home directory or any directory you prefer (e.g., `/home/riza/`).
Unpack the tarball and set the environment variables as follows:
tar zxvf jdk-24_linux-x64_bin.tar.gz
nano ~/.bashrc
Add the following lines at the end of your ~/.bashrc file (edit the code according to the location of the file on your machine)
export JAVA_HOME='/home/riza/jdk-24'
export PATH=$JAVA_HOME/bin:$PATH
PICARD Installation
Download the latest jar file here or use:
wget https://github.com/broadinstitute/picard/releases/download/3.3.0/picard.jar
Place the file in your home directory or any directory you prefer (e.g., `/home/riza/`).
Set the environment variable:
nano ~/.bashrc
Add the following lines at the end of your ~/.bashrc file (edit the code according to the location of the file on your machine)
export PICARD='/home/riza/picard.jar'
Verify installation:
java -jar $PICARD
GATK4 Installation
Download the latest file here or use:
wget https://github.com/broadinstitute/gatk/releases/download/4.6.1.0/gatk-4.6.1.0.zip
Place the file in your home directory or any directory you prefer (e.g., `/home/riza/`).
Set the environment variable:
nano ~/.bashrc
Add the following lines at the end of your ~/.bashrc file (edit the code according to the location of the file on your machine)
export GATK='/home/riza/gatk-4.6.1.0/gatk-package-4.6.1.0-local.jar'
Verify installation:
java -jar $GATK
MacOS-specific
- Homebrew
- Follow installation instructions at the homepage of Homebrew
- Or on the Terminal app
- /bin/bash -c "$(curl -fsSL https://raw.githubusercontent.com/Homebrew/install/HEAD/install.sh)"
- Run these two commands in your terminal to add Homebrew to your PATH:
- (echo; echo 'eval "$(/usr/local /bin/brew shellenv)"')
- eval "$(/usr/local/bin/brew shellenv)"
- In the terminal app
- brew install bwa
- brew install fastp
- brew install fasqc
- brew install samtools
- brew install bcftools
- brew install picard-tools


